Multiplexed single-molecule spectroscopy for ultrasensitive biomarker sensing
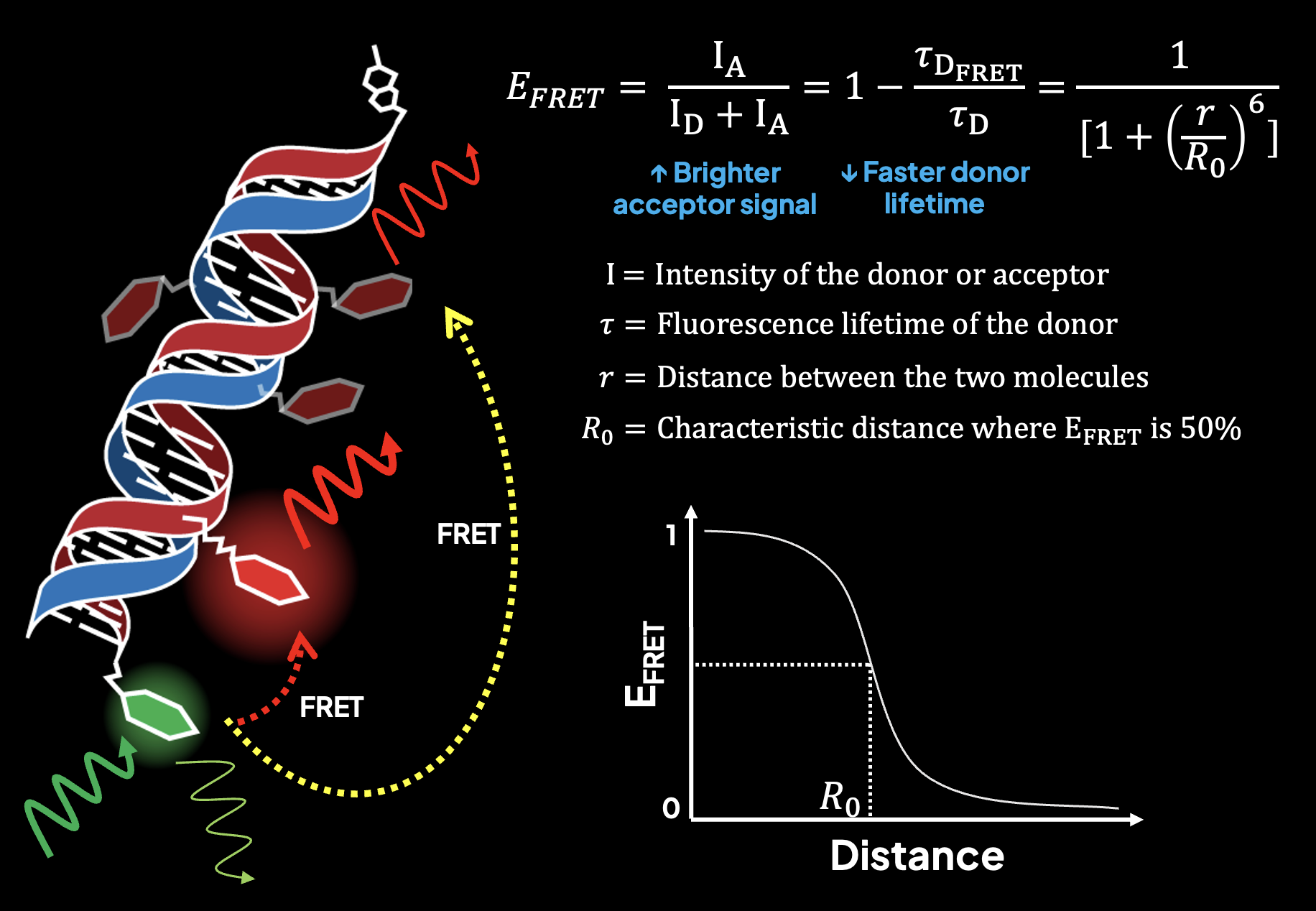
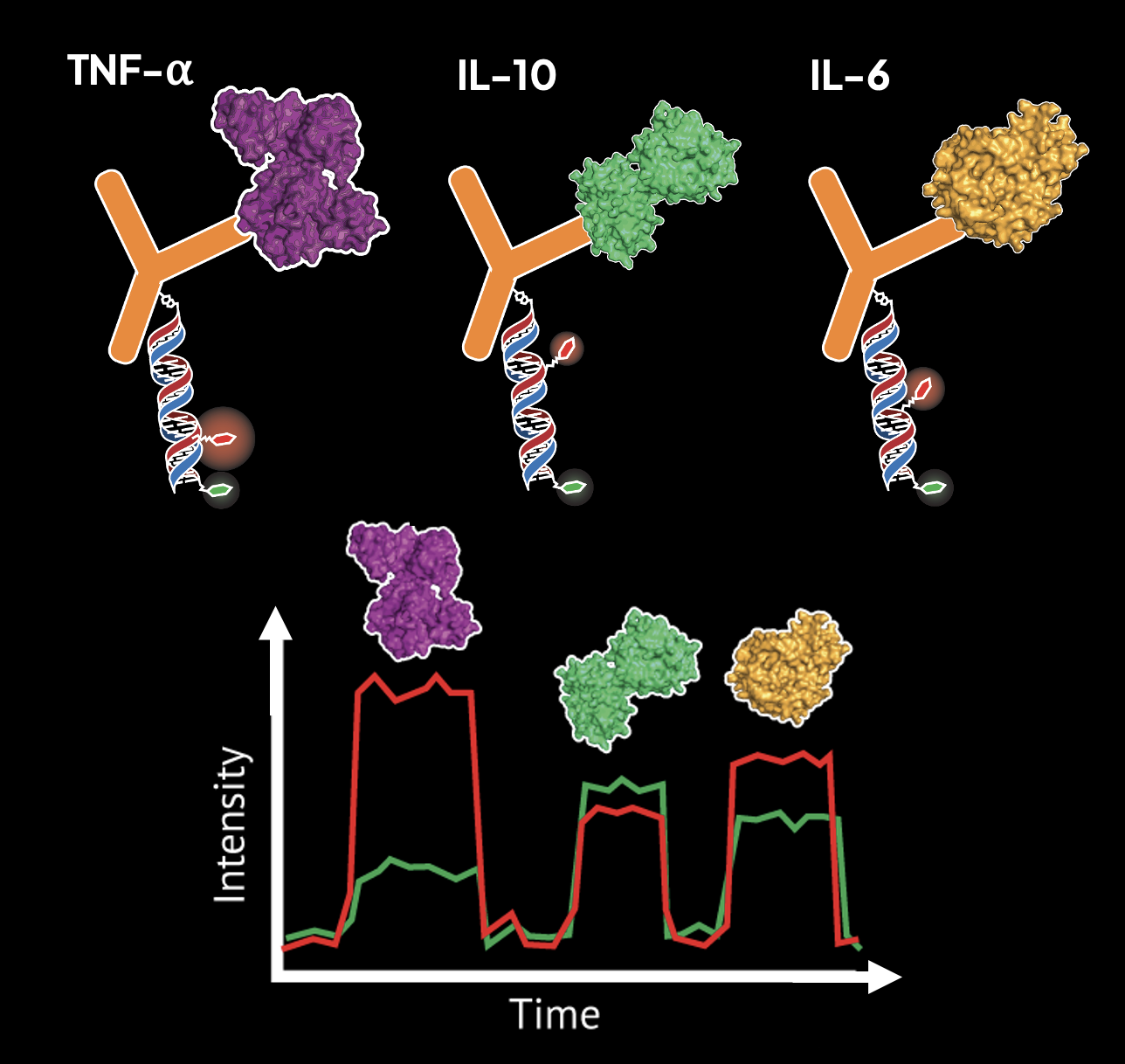
Monitoring many biomolecules and their interactions is crucial for understanding biological complexity, yet many of the signaling molecules informing the early-phase dynamics of a response are present at amounts far below the sensitivity of conventional biomolecular profiling assays. In order to approach the very limits of detection, profiling these molecules at the single-molecule level is needed. Single-molecule fluorescence spectroscopy enables information-rich measurements of biomolecules present at ultra-low abundance. However, factors such as spectral crowding and low signal-to-noise have severely limited multiplexing capabilities in these regimes, leaving many biological systems whose molecular logic remains poorly understood because we lack the tools to interrogate them. Previously, we developed FRETfluors, novel DNA-based FRET constructs that possess differing emission signatures that enable high-order multiplexed sensing at the single-molecule level with only two emission channels. We are working towards an advancement in single-molecule biomarker sensing by pairing FRETfluors with affinity-based sensing and data-driven inference of biomolecular identity. FRETfluors are covalently conjugated to antibodies to report single-molecule binding events to a panel of biomarkers, after which the Anti-Brownian Electrokinetic (ABEL) trap is used to identify spectroscopic fingerprints for downstream analysis. To further unlock the information latent in photon-by-photon data such as post-translational modifications and binding states, we are also developing a machine learning pipeline to learn classifying features and extract hidden observables that report more information on the biophysical characteristics of the biomolecules usually inaccessible with sequence-based readouts. In turn, our platform will enable multiplexed biomolecular profiling studies of complex and input-scarce systems at the single-molecule level that can help us understand complex signaling responses with very high spatiotemporal resolution.
Who is involved: Ryan Febryanto (jointly with Allison Squires’ lab)