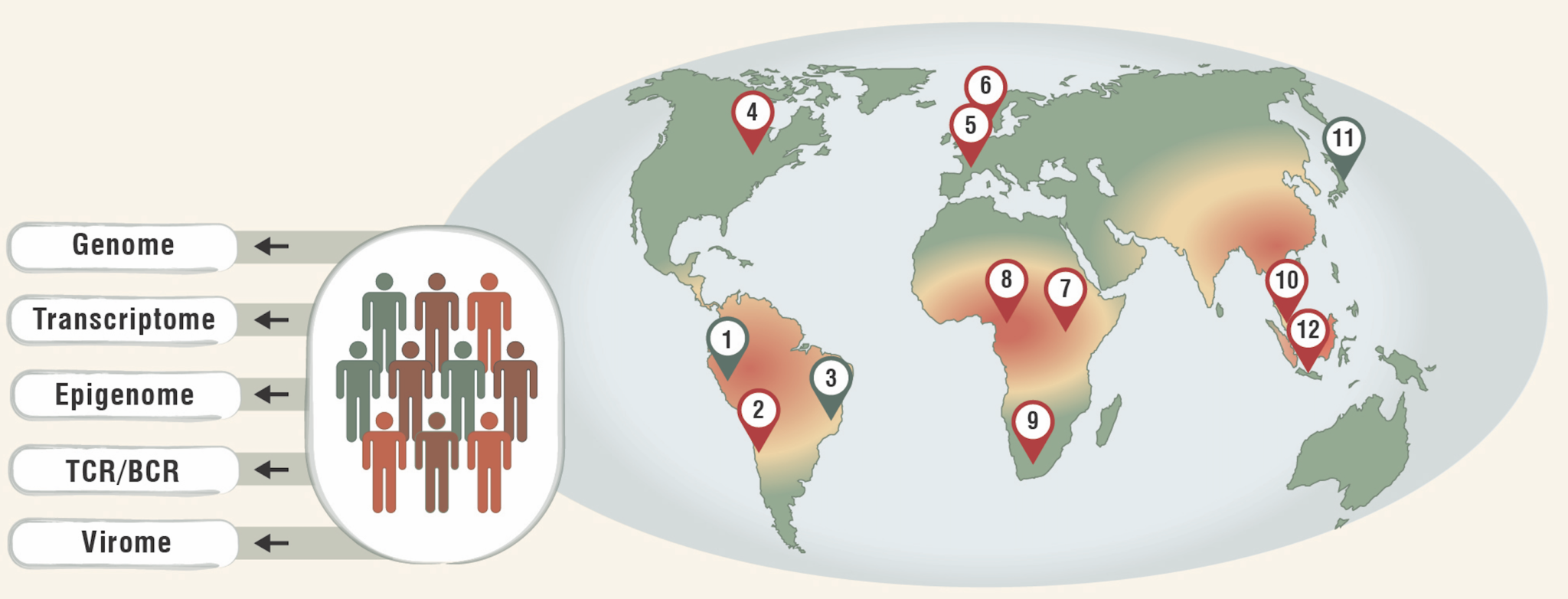
Immune Variation Across Diverse Human Populations
Individual differences in susceptibility to infections, disease severity, and therapeutic response result from a complex interplay of genetic and environmental factors. However, progress in defining the drivers of immune variation remains limited because most genomic studies focus on European populations, restricting the transferability of genetic insights. Furthermore, these studies lack the environmental diversity needed to understand the implications for immunity of industrialization-associated diets, physical activity patterns, and toxin exposure landscapes. To bridge this gap, we aim to profile the immune system in populations with varied genetic ancestries, environments, and lifestyles. We have partnered with global collaborators to collect and analyze peripheral blood mononuclear cells (PBMCs) from genotyped individuals across myriad populations, including underrepresented and Indigenous populations. We will comprehensively characterize chromatin accessibility and transcriptional regulation patterns in all major immune cell types at single-cell resolution to elucidate epigenetic mechanisms that may play a role in the regulation of immune responses. Our analysis of these data will identify epigenetic and transcriptomic factors, along with, when relevant, their genetic associations, that contribute to the biological basis of variation in susceptibility to disease. In turn, these results will enable better prediction of the effects of genetic and environmental contributors on immune responses and lend insights into health disparities in minority, Indigenous, and other underrepresented groups.
Who is involved: Akansha Gupta (jointly with Luis Barreiro’s lab)